
Researchers mapped genetic blueprints for 51 species including cats, dolphins, kangaroos, penguins, sharks, and turtles, a discovery that deepens our understanding of evolution and the links between humans and animals.
“Being able to access that genetic information will have huge implications for understanding human health and evolution,” said lead author Michael Schatz, a Bloomberg Distinguished Professor of computer science and biology at Johns Hopkins University. “A lot of work on drug compounds starts in mice and other animal models, so understanding their genomes and the genomes of other animals directly benefits us.”
The team, working with the Vertebrate Genomes Project, sequenced the genomes of 51 vertebrate species, prioritizing those that are useful models for understanding human evolution. The researchers developed novel algorithms and computer software that cut the sequencing time from months—or decades in the case of the human genome—to a matter of days.
The findings, “Scalable, accessible and reproducible reference genome assembly and evaluation in Galaxy,” are newly published in the journal Nature Biotechnology.
Mammals, a subset of vertebrates that includes primates, dogs, cats, mice, and humans, share 50% to 99% of the same DNA and nearly all the genes from a common ancestor that lived roughly 200 million years ago. By comparing the complete genomes of these species, researchers can start to identify when and where DNA sequences diverged and the implications of those differences for humans. But, researchers say, this work has been limited by the number and quality of vertebrate genomes available, which has focused on a few key species.
Vertebrate genomes are billions of characters long, too long for any gene sequencing technology to read in one complete pass. Researchers must rely on tools that break down the genome into smaller, easier to read segments. Computer programs then take those segments and determine how they fit together, like pieces of a jigsaw puzzle.
But traditional technology was not able to finish the puzzle.
“Have you ever done a massive jigsaw puzzle where at some point all that’s left is blue sky, and you don’t think you’ll ever be able to fit the right pieces together? The old software would basically give up on these hard parts of the genome. That’s the problem with genome assembly,” Schatz said. “Our new program, using the latest sequencing data and the latest assembly algorithms, knows how to work through those parts to get a more complete picture.”
To test their technology, researchers mapped the genome of the zebra finch, a songbird that had already been sequenced to study brain development. The new technology was far better at reassembling segments of the genome, creating a more accurate and complete map.
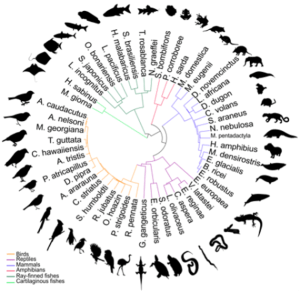
Researchers selected these 51 animals for gene sequencing, deepening our understanding of how they’re related to each other. Credit: Delphine Lariviere, Penn State University.
The open-source software is available online via Galaxy, a web-based platform, based at Johns Hopkins and Penn State, that offers scientific software for free to the public and supports half a million scientists and educators worldwide.
“In the past, only a handful of elite research groups would have had access to the resources needed to assemble these genomes. Now, anyone on the planet with access to the internet can visit the website and, with a few clicks of the button, run multiple scientific tools,” said Alex Ostrovsky, a Johns Hopkins software engineer on the Galaxy team who was responsible for making the tools easy to use for noncoders.
The team will continue working with the Vertebrate Genomes Project to sequence the genomes of at least one species across all 275 vertebrate orders.
“In some ways, we’re building an evolutionary time machine,” Schatz said. “We can trace how vertebrates evolved over time and eventually gave rise to genes and sequences that are uniquely found in humans.
“Having the genes of our evolutionary cousins mapped out will help us better understand ourselves.”
This work was funded by NIH Grants U41 HG006620, U24 HG010263, U24 CA231877 and U01CA253481, along with NSF Grants 1661497, 1758800, and 2216612. The work was supported in part by The Human Frontier Science Program RGP0025/2021; the Swiss National Science Foundation grants 202669 and 198691; the Swiss State Secretariat for Education, Research and Innovation grant 22.00173; and Horizon Europe under the Biodiversity, Circular Economy and Environment program (REA.B.3, BGE101059492). Usegalaxy.eu is supported by the German Federal Ministry of Education and Research grants 031L0101C and de.NBI-epi.
This work was performed in collaboration with researchers at Pennsylvania State University, Rockefeller University, and several other institutions. Computational resources were provided by the Advanced Cyberinfrastructure Coordination Ecosystem (ACCESS-CI), the Texas Advanced Computing Center, the JetStream2 scientific cloud, and the Rockfish data center at Johns Hopkins University.