Clinical neuroscience is a field with all the difficulties that come from high dimensional datasets, and none of the advantages that fuel modern-day breakthroughs in domains such as computer vision and natural language processing. It is a field of small sample sizes, complex phenomena, massive patient variability, and most importantly, an arguable lack of ground truth information. The NSA Lab tackles these challenges via a two-pronged approach. On the application front, we identify key knowledge gaps that can advance either our understanding or management of a disorder. On the methodological side, we combine hypothesis-driven insights about the modalities, often from our collaborators, with data-driven learning techniques to address these gaps.
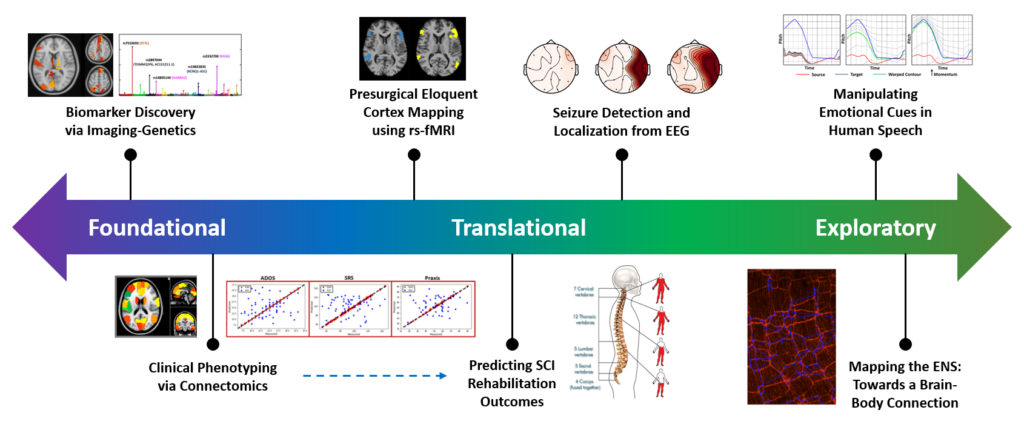
Our current projects span the gamut from foundational computational neuroscience to translational applications of functional data to exploratory directions for probing neural circuitry. This multi-pronged approach allow us to generalize the algorithms across datasets and applications. Additionally, our unique modeling strategies can inspire solutions for complex engineering problems outside of the medical realm.