EPViz (EEG Prediction Visualizer)
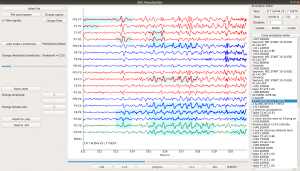 Scalp EEG is one of the most popular noninvasive modalities for studying real-time neural phenomena. While traditional EEG studies have focused on identifying group-level statistical effects, the rise of machine learning has prompted a shift in the community towards spatio-temporal predictive analyses. We have developed the EEG Prediction Visualizer (EPViz) to aid researchers in developing, validating, and reporting their predictive modeling outputs. EPViz is a lightweight and standalone software package developed in Python. Beyond viewing and manipulating the EEG data, EPViz allows researchers to load a PyTorch deep learning model, apply it to the EEG, and overlay the output channel-wise or subject-level temporal predictions on top of the original time series. These results can be saved as high-resolution images for use in manuscripts and presentations. There is also a command-line option for batch processing. EPViz also provides valuable tools for clinician-scientists, including spectrum visualization, computation of basic statistics, data anonymization, and annotation editing.
Scalp EEG is one of the most popular noninvasive modalities for studying real-time neural phenomena. While traditional EEG studies have focused on identifying group-level statistical effects, the rise of machine learning has prompted a shift in the community towards spatio-temporal predictive analyses. We have developed the EEG Prediction Visualizer (EPViz) to aid researchers in developing, validating, and reporting their predictive modeling outputs. EPViz is a lightweight and standalone software package developed in Python. Beyond viewing and manipulating the EEG data, EPViz allows researchers to load a PyTorch deep learning model, apply it to the EEG, and overlay the output channel-wise or subject-level temporal predictions on top of the original time series. These results can be saved as high-resolution images for use in manuscripts and presentations. There is also a command-line option for batch processing. EPViz also provides valuable tools for clinician-scientists, including spectrum visualization, computation of basic statistics, data anonymization, and annotation editing.
EPViz can be installed in three ways: (1) cloning our GitHub repository to access the latest version, (2) through PyPI, and (3) as a standalone prepackaged application for MacOS and Windows. [github][pypi][Mac App][Windows App].
Please visit our EPViz page for the complete user guide.
EDF Anonymizer
 We have also provided just the anonymization tool in EPViz as a standalone module. This tool allows the user to alter the EDF header fields and provides default settings for scrubbing patient IDs and time stamps. [github][Windows App][MAC App]
We have also provided just the anonymization tool in EPViz as a standalone module. This tool allows the user to alter the EDF header fields and provides default settings for scrubbing patient IDs and time stamps. [github][Windows App][MAC App]
Epilepsy
- N. Nandakumar, D. Hsu, R. Ahmed, A. Venkataraman. DeepSOZ: A Graph Convolutional Network for Automated Seizure Onset Zone Localization from Resting-State fMRI Connectivity. Under Revision for IEEE Transactions on Biomedical Engineering, 2022. [paper][github]
Predictive Connectomics
- N.S. D’Souza, M.B. Nebel, D. Crocetti, N. Wymbs, J. Robinson, S. Mostofsky, A. Venkataraman. A Matrix Auto-encoder Framework to Align the Functional and Structural Connectivity Manifolds as Guided by Behavioral Phenotypes. In Proc. MICCAI: Medical Image Computing and Computer Assisted Intervention, LNCS (to appear), 2021. [paper][github]
- N.S. D’Souza, M.B. Nebel, D. Crocetti, N. Wymbs, J. Robinson, S. Mostofsky, A. Venkataraman. M-GCN: A Multimodal Graph Convolutional Network to Integrate Functional and Structural Connectomics Data to Predict Multidimensional Phenotypic Characterizations. In Proc. MIDL: Medical Imaging with Deep Learning, MLR:1-12, 2021. [paper][github]
- N.S. D’Souza, M.B. Nebel, D. Crocetti, N. Wymbs, J. Robinson, S. Mostofsky, A. Venkataraman. A Deep-Generative Hybrid Model to Integrate Multimodal and Dynamic Connectivity for Predicting Spectrum-Level Deficits in Autism. In Proc. MICCAI: Medical Image Computing and Computer Assisted Intervention, LNCS 12267:437-447, 2020. [paper][github]
- N.S. D’Souza, N. Wymbs, M.B. Nebel, S. Mostofsky, A. Venkataraman. A Joint Network Optimization Framework to Predict Clinical Severity from Resting State fMRI Data. NeuroImage, 206:116314, 2020. [paper][github]
Imaging-Genetics
- A Generative Discriminative Framework that Integrates Imaging, Genetic, and Diagnosis into Coupled Low Dimensional Space [paper][github]
Preoperative Mapping with Resting-State fMRI
- A Multi-Scale Spatial and Temporal Attention Network on Dynamic Connectivity to Localize The Eloquent Cortex in Brain Tumor Patients [paper][github]
- A Multi-Task Deep Learning Framework to Localize the Eloquent Cortex in Brain Tumor Patients using Both Static and Dynamic Functional Conn [paper][github]
Emotional Speech
- Sample, Attend and Morph: A Deep-Bayesian Framework for Adaptive Speech Duration Modification [paper][github]
- Non-parallel Emotion Conversion using a Deep-Generative Hybrid Network and an Adversarial Pair Discriminator [paper][github]
- Multi-Speaker Emotion Conversion via Latent Variable Regularization and a Chained Encoder-Decoder-Predictor Network [paper][github]
- A Diffeomorphic Flow-based Variational Framework for Multi-speaker Emotion Conversion [paper][github][sample audio]